Human leukemia K562 cells: induction to erythroid differentiation by
Guanosine Và Guanine Nucleotides kháng K562
Guanine
From Wikipedia, the free encyclopedia
 | |||
| Names | |||
|---|---|---|---|
| IUPAC name
2-amino-9H-purin-6(1H)-one
| |||
| Other names
1,9-dihydro-6H-purin-6-one,
2-amino-6-hydroxypurine, 2-aminohypoxanthine, Guanine | |||
| Identifiers | |||
3D model (JSmol)
| |||
| ChEBI | |||
| ChemSpider | |||
| DrugBank | |||
| ECHA InfoCard | 100.000.727 | ||
| KEGG | |||
| RTECS number | MF8260000 | ||
| UNII | |||
| Properties | |||
| C5H5N5O | |||
| Molar mass | 151.13 g/mol | ||
| Appearance | White amorphous solid. | ||
| Density | 2.200 g/cm3 (calculated) | ||
| Melting point | 360 °C (680 °F; 633 K) decomposes | ||
| Boiling point | Sublimes | ||
| Insoluble. | |||
| Acidity (pKa) | 3.3 (amide), 9.2 (secondary), 12.3 (primary)[1] | ||
| Hazards | |||
| Main hazards | Irritant | ||
| NFPA 704 | |||
| Flash point | Non-flammable | ||
| Related compounds | |||
Related compounds
| Cytosine; Adenine; Thymine; Uracil | ||
Except where otherwise noted, data are given for materials in their standard state (at 25 °C [77 °F], 100 kPa).
| |||
| Infobox references | |||
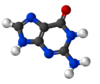
Guanine (/ˈɡwɑːnᵻn/; or G, Gua) is one of the four main nucleobases found in the nucleic acids DNA and RNA, the others being adenine, cytosine, and thymine (uracil in RNA). In DNA, guanine is paired with cytosine. The guanine nucleoside is called guanosine.
With the formula C5H5N5O, guanine is a derivative of purine, consisting of a fused pyrimidine-imidazole ring system with conjugated double bonds. Being unsaturated, the bicyclic molecule is planar.
Contents
[hide]Properties[edit]
Guanine, along with adenine and cytosine, is present in both DNA and RNA, whereas thymine is usually seen only in DNA, and uracil only in RNA. Guanine has two tautomeric forms, the major keto form (see figures) and rare enol form. It binds to cytosine through three hydrogen bonds. In cytosine, the amino group acts as the hydrogen bond donor and the C-2 carbonyl and the N-3 amine as the hydrogen-bond acceptors. Guanine has the C-6 carbonyl group that acts as the hydrogen bond acceptor, while a group at N-1 and the amino group at C-2 act as the hydrogen bond donors.
 |  |
Guanine can be hydrolyzed with strong acid to glycine, ammonia, carbon dioxide, and carbon monoxide. First, guanine gets deaminated to become xanthine.[2] Guanine oxidizes more readily than adenine, the other purine-derivative base in DNA. Its high melting point of 350 °C reflects the intermolecular hydrogen bonding between the oxo and amino groups in the molecules in the crystal. Because of this intermolecular bonding, guanine is relatively insoluble in water, but it is soluble in dilute acids and bases.
History[edit]
The first isolation of guanine was reported in 1844 by the German chemist Julius Bodo Unger (1819–1885), who obtained it from the excreta of sea birds, which is known as guano and which was used as a source of fertilizer; guanine was named in 1846.[3] Between 1882 and 1906, Fischer determined the structure and also showed that uric acid can be converted to guanine.[4]
Syntheses[edit]
Trace amounts of guanine form by the polymerization of ammonium cyanide (NH
4CN). Two experiments conducted by Levy et al. showed that heating 10 mol·L−1 NH
4CN at 80 °C for 24 hours gave a yield of 0.0007%, while using 0.1 mol·L−1NH
4CN frozen at −20 °C for 25 years gave a 0.0035% yield. These results indicate guanine could arise in frozen regions of the primitive earth. In 1984, Yuasa reported a 0.00017% yield of guanine after the electrical discharge of NH
3, CH
4, C
2H
6, and 50 mL of water, followed by a subsequent acid hydrolysis. However, it is unknown whether the presence of guanine was not simply a resultant contaminant of the reaction.[5]
4CN). Two experiments conducted by Levy et al. showed that heating 10 mol·L−1 NH
4CN at 80 °C for 24 hours gave a yield of 0.0007%, while using 0.1 mol·L−1NH
4CN frozen at −20 °C for 25 years gave a 0.0035% yield. These results indicate guanine could arise in frozen regions of the primitive earth. In 1984, Yuasa reported a 0.00017% yield of guanine after the electrical discharge of NH
3, CH
4, C
2H
6, and 50 mL of water, followed by a subsequent acid hydrolysis. However, it is unknown whether the presence of guanine was not simply a resultant contaminant of the reaction.[5]
- 10NH3 + 2CH4 + 4C2H6 + 2H2O → 2C5H8N5O (guanine) + 25H2
A Fischer-Tropsch synthesis can also be used to form guanine, along with adenine, uracil, and thymine. Heating an equimolar gas mixture of CO, H2, and NH3 to 700 °C for 15 to 24 minutes, followed by quick cooling and then sustained reheating to 100 to 200 °C for 16 to 44 hours with an alumina catalyst, yielded guanine and uracil:
- 10CO + H2 + 10NH3 → 2C5H8N5O (guanine) + 8H2O
Another possible abiotic route was explored by quenching a 90% N2–10%CO–H2O gas mixture high-temperature plasma.[6]
Traube's synthesis involves heating 2,4,5-triamino-1,6-dihydro-6-oxypyrimidine (as the sulfate) with formic acid for several hours. 

Other occurrences/ biological uses[edit]
The word guanine derives from the Spanish loanword guano ("bird/bat droppings"), which itself is from the Quechua word wanu, meaning "dung". As the Oxford English Dictionary notes, guanine is "A white amorphous substance obtained abundantly from guano, forming a constituent of the excrement of birds".[7]
In 1656 in Paris, a Mr. Jaquin extracted from the scales of the fish Alburnus alburnus so-called "pearl essence",[8] which is crystalline guanine.[9] In the cosmetics industry, crystalline guanine is used as an additive to various products (e.g., shampoos), where it provides a pearly iridescent effect. It is also used in metallic paints and simulated pearls and plastics. It provides shimmering luster to eye shadow and nail polish. Facial treatments using the droppings, or guano, from Japanese nightingales have been used in Japan and elsewhere, reportedly because the guanine in the droppings produces a clear, "bright" skin tone[10] that users desire. Guanine crystals are rhombic platelets composed of multiple transparent layers, but they have a high index of refraction that partially reflects and transmits light from layer to layer, thus producing a pearly luster. It can be applied by spray, painting, or dipping. It may irritate the eyes. Its alternatives are mica, faux pearl (from ground shells),[11] and aluminium and bronze particles.
Spiders and scorpions convert ammonia, as a product of protein metabolism in the cells, to guanine, as it can be excreted with minimal water loss.
Guanine is found in the skin of many fish (e.g., the sturgeon).[12] It is also present in the reflective deposits of the eyes of deep-sea fish and some reptiles, such as crocodiles.[12]
On 8 August 2011, a report, based on NASA studies with meteorites found on Earth, was published suggesting building blocks of DNA and RNA (guanine, adenine and related organic molecules) may have been formed extraterrestrially in outer space.[13][14][15]
Guanosine
From Wikipedia, the free encyclopedia
| This article needs additional citations for verification. (January 2015) (Learn how and when to remove this template message) |
 | |
 | |
| Names | |
|---|---|
| IUPAC name
2-Amino-9-[3,4-dihydroxy-5-(hydroxymethyl)oxolan-2-yl]-3H-purin-6-one
| |
| Other names
Guanine riboside
| |
| Identifiers | |
3D model (JSmol)
| |
| ChEBI | |
| ChemSpider | |
| DrugBank | |
| ECHA InfoCard | 100.003.844 |
| KEGG | |
| MeSH | Guanosine |
PubChem CID
| |
| UNII | |
| Properties | |
| C10H13N5O5 | |
| Molar mass | 283.241 |
| -149.1·10−6 cm3/mol | |
Except where otherwise noted, data are given for materials in their standard state (at 25 °C [77 °F], 100 kPa).
| |
| Infobox references | |
Guanosine is a purine nucleoside comprising guanine attached to a ribose (ribofuranose) ring via a β-N9-glycosidic bond. Guanosine can be phosphorylated to become guanosine monophosphate (GMP), cyclic guanosine monophosphate(cGMP), guanosine diphosphate (GDP), and guanosine triphosphate (GTP). These forms play important roles in various biochemical processes such as synthesis of nucleic acids and proteins, photosynthesis, muscle contraction, and intracellular signal transduction (cGMP). When guanine is attached by its N9 nitrogen to the C1 carbon of a deoxyribosering it is known as deoxyguanosine.
The antiviral drug aciclovir, often used in herpes treatment, and the anti-HIV drug abacavir, are structurally similar to guanosine.
Guanosine is required for an RNA splicing reaction in mRNA, when a "self-splicing" intron removes itself from the mRNA message by cutting at both ends, re-ligating, and leaving just the exons on either side to be translated into protein.[1]
Guanosine was used to make Regadenoson.
References[edit]
Nucleotide
From Wikipedia, the free encyclopedia
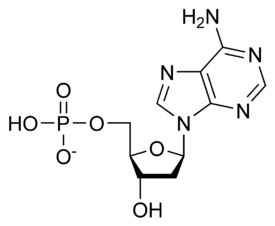
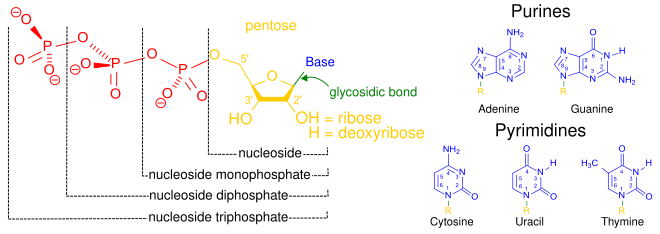
Nucleotides are organic molecules that serve as the monomer units for forming the nucleic acid polymers DNA(deoxyribonucleic acid) and RNA (ribonucleic acid), both of which are essential biomolecules in all life-forms on Earth. Nucleotides are the building blocks of nucleic acids; they are composed of three subunit molecules: a nitrogenous base, a five-carbon sugar (ribose or deoxyribose), and at least one phosphate group. They are also known as phosphate nucleotides.
A nucleoside is a nitrogenous base and a 5-carbon sugar. Thus a nucleoside plus a phosphate group yields a nucleotide.
Nucleotides also play a central role in life-form metabolism at the fundamental, cellular level. They carry packets of chemical energy—in the form of the nucleoside triphosphates ATP, GTP, CTP and UTP—throughout the cell to the many cellular functions that demand energy, which include synthesizing amino acids, proteins and cell membranes and parts; moving the cell and moving cell parts, both internally and intercellularly; dividing the cell, etc.[1] In addition, nucleotides participate in cell signaling (cGMP and cAMP), and are incorporated into important cofactors of enzymatic reactions (e.g. coenzyme A, FAD, FMN, NAD, and NADP+).
In experimental biochemistry, nucleotides can be radiolabeled with radionuclides to yield radionucleotides.
Contents
[hide]Structure[edit]
A nucleotide is composed of three distinctive chemical sub-units: a five-carbon sugar molecule, a nitrogenous base—which two together are called a nucleoside—and one phosphate group. With all three joined, a nucleotide is also termed a "nucleoside monophosphate". The chemistry sources ACS Style Guide[2] and IUPAC Gold Book[3] prescribe that a nucleotide should contain only one phosphate group, but common usage in molecular biology textbooks often extends the definition to include molecules with two, or with three, phosphates.[1][4][5][6] Thus, the terms "nucleoside diphosphate" or "nucleoside triphosphate" may also indicate nucleotides.
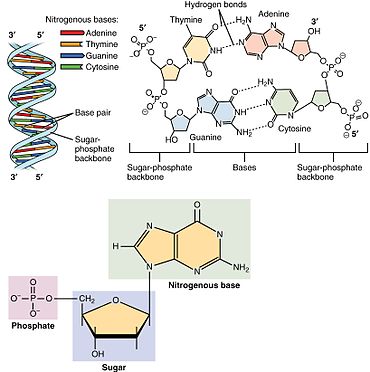
Nucleotides contain either a purine or a pyrimidine base—i.e., the nitrogenous base molecule, also known as a nucleobase—and are termed ribonucleotides if the sugar is ribose, or deoxyribonucleotides if the sugar is deoxyribose. Individual phosphate molecules repetitively connect the sugar-ringmolecules in two adjacent nucleotide monomers, thereby connecting the nucleotide monomers of a nucleic acid end-to-end into a long chain. These chain-joins of sugar and phosphate molecules create a 'backbone' strand for a single- or double helix. In any one strand, the chemical orientation (directionality) of the chain-joins runs from the 5'-end to the 3'-end (read: 5 prime-end to 3 prime-end)—referring to the five carbon sites on sugar molecules in adjacent nucleotides. In a double helix, the two strands are oriented in opposite directions, which permits base pairing and complementaritybetween the base-pairs, all which is essential for replicating or transcribing the encoded information found in DNA.
Unlike in nucleic acid nucleotides, singular cyclic nucleotides are formed when the phosphate group is bound twice to the same sugar molecule, i.e., at the corners of the sugar hydroxyl groups.[1] These individual nucleotides function in cell metabolism rather than the nucleic acid structures of long-chain molecules.
Nucleic acids then are polymeric macromolecules assembled from nucleotides, the monomer-units of nucleic acids. The purine bases adenine and guanine and pyrimidine base cytosine occur in both DNA and RNA, while the pyrimidine bases thymine (in DNA) and uracil (in RNA) in just one. Adenine forms a base pair with thymine with two hydrogen bonds, while guanine pairs with cytosine with three hydrogen bonds.
Synthesis[edit]
In vivo, nucleotides can be synthesized de novo or recycled through salvage pathways.[7] The components used in de novo nucleotide synthesis are derived from biosynthetic precursors of carbohydrate and amino acid metabolism, and from ammonia and carbon dioxide. The liver is the major organ of de novo synthesis of all four nucleotides. De novo synthesis of pyrimidines and purines follows two different pathways. Pyrimidines are synthesized first from aspartate and carbamoyl-phosphate in the cytoplasm to the common precursor ring structure orotic acid, onto which a phosphorylated ribosyl unit is covalently linked. Purines, however, are first synthesized from the sugar template onto which the ring synthesis occurs. For reference, the syntheses of the purine and pyrimidine nucleotides are carried out by several enzymes in the cytoplasm of the cell, not within a specific organelle. Nucleotides undergo breakdown such that useful parts can be reused in synthesis reactions to create new nucleotides.
In vitro, protecting groups may be used during laboratory production of nucleotides. A purified nucleoside is protected to create a phosphoramidite, which can then be used to obtain analogues not found in nature and/or to synthesize an oligonucleotide.
Pyrimidine ribonucleotide synthesis[edit]
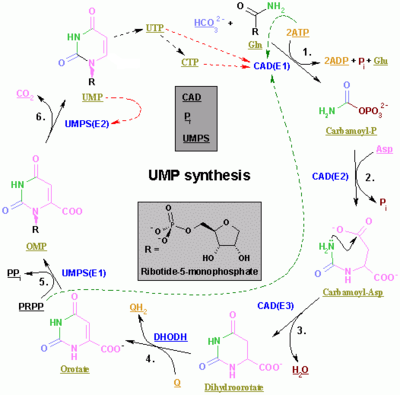
The synthesis of the pyrimidines CTP and UTP occurs in the cytoplasm and starts with the formation of carbamoyl phosphate from glutamine and CO2. Next, aspartate carbamoyltransferasecatalyzes a condensation reaction between aspartate and carbamoyl phosphate to form carbamoyl aspartic acid, which is cyclized into 4,5-dihydroorotic acid by dihydroorotase. The latter is converted to orotate by dihydroorotate oxidase. The net reaction is:
- (S)-Dihydroorotate + O2 → Orotate + H2O2
Orotate is covalently linked with a phosphorylated ribosyl unit. The covalent linkage between the ribose and pyrimidine occurs at position C1[8] of the ribose unit, which contains a pyrophosphate, and N1 of the pyrimidine ring. Orotate phosphoribosyltransferase (PRPP transferase) catalyzes the net reaction yielding orotidine monophosphate (OMP):
- Orotate + 5-Phospho-α-D-ribose 1-diphosphate (PRPP) → Orotidine 5'-phosphate + Pyrophosphate
Orotidine 5'-monophosphate is decarboxylated by orotidine-5'-phosphate decarboxylase to form uridine monophosphate (UMP). PRPP transferase catalyzes both the ribosylation and decarboxylation reactions, forming UMP from orotic acid in the presence of PRPP. It is from UMP that other pyrimidine nucleotides are derived. UMP is phosphorylated by two kinases to uridine triphosphate (UTP) via two sequential reactions with ATP. First the diphosphate form UDP is produced, which in turn is phosphorylated to UTP. Both steps are fueled by ATP hydrolysis:
- ATP + UMP → ADP + UDP
- UDP + ATP → UTP + ADP
CTP is subsequently formed by amination of UTP by the catalytic activity of CTP synthetase. Glutamine is the NH3 donor and the reaction is fueled by ATP hydrolysis, too:
- UTP + Glutamine + ATP + H2O → CTP + ADP + Pi
Cytidine monophosphate (CMP) is derived from cytidine triphosphate (CTP) with subsequent loss of two phosphates.[9] [10]
Purine ribonucleotide synthesis[edit]
The atoms that are used to build the purine nucleotides come from a variety of sources:
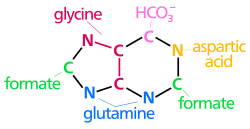 | The biosyntheticorigins of purine ring atoms N1 arises from the amine group of Asp C2 and C8 originate from formate N3 and N9 are contributed by the amide group of Gln C4, C5 and N7 are derived from Gly C6 comes from HCO3− (CO2) |
The de novo synthesis of purine nucleotides by which these precursors are incorporated into the purine ring proceeds by a 10-step pathway to the branch-point intermediate IMP, the nucleotide of the base hypoxanthine. AMP and GMP are subsequently synthesized from this intermediate via separate, two-step pathways. Thus, purine moieties are initially formed as part of the ribonucleotides rather than as free bases.
Six enzymes take part in IMP synthesis. Three of them are multifunctional:
The pathway starts with the formation of PRPP. PRPS1 is the enzyme that activates R5P, which is formed primarily by the pentose phosphate pathway, to PRPP by reacting it with ATP. The reaction is unusual in that a pyrophosphoryl group is directly transferred from ATP to C1 of R5P and that the product has the α configuration about C1. This reaction is also shared with the pathways for the synthesis of Trp, His, and the pyrimidine nucleotides. Being on a major metabolic crossroad and requiring much energy, this reaction is highly regulated.
In the first reaction unique to purine nucleotide biosynthesis, PPAT catalyzes the displacement of PRPP's pyrophosphate group (PPi) by an amide nitrogen donated from either glutamine (N), glycine (N&C), aspartate (N), folic acid (C1), or CO2. This is the committed step in purine synthesis. The reaction occurs with the inversion of configuration about ribose C1, thereby forming β-5-phosphorybosylamine (5-PRA) and establishing the anomeric form of the future nucleotide.
Next, a glycine is incorporated fueled by ATP hydrolysis and the carboxyl group forms an amine bond to the NH2 previously introduced. A one-carbon unit from folic acid coenzyme N10-formyl-THF is then added to the amino group of the substituted glycine followed by the closure of the imidazole ring. Next, a second NH2 group is transferred from a glutamine to the first carbon of the glycine unit. A carboxylation of the second carbon of the glycin unit is concomittantly added. This new carbon is modified by the additional of a third NH2 unit, this time transferred from an aspartate residue. Finally, a second one-carbon unit from formyl-THF is added to the nitrogen group and the ring covalently closed to form the common purine precursor inosine monophosphate (IMP).
Inosine monophosphate is converted to adenosine monophosphate in two steps. First, GTP hydrolysis fuels the addition of aspartate to IMP by adenylosuccinate synthase, substituting the carbonyl oxygen for a nitrogen and forming the intermediate adenylosuccinate. Fumarate is then cleaved off forming adenosine monophosphate. This step is catalyzed by adenylosuccinate lyase.
Inosine monophosphate is converted to guanosine monophosphate by the oxidation of IMP forming xanthylate, followed by the insertion of an amino group at C2. NAD+is the electron acceptor in the oxidation reaction. The amide group transfer from glutamine is fueled by ATP hydrolysis.
Pyrimidine and purine degradation[edit]
In humans, pyrimidine rings (C, T, U) can be degraded completely to CO2 and NH3 (urea excretion). That having been said, purine rings (G, A) cannot. Instead they are degraded to the metabolically inert uric acid which is then excreted from the body. Uric acid is formed when GMP is split into the base guanine and ribose. Guanine is deaminated to xanthine which in turn is oxidized to uric acid. This last reaction is irreversible. Similarly, uric acid can be formed when AMP is deaminated to IMP from which the ribose unit is removed to form hypoxanthine. Hypoxanthine is oxidized to xanthine and finally to uric acid. Instead of uric acid secretion, guanine and IMP can be used for recycling purposes and nucleic acid synthesis in the presence of PRPP and aspartate (NH3 donor).
Unnatural base pair (UBP)[edit]
An unnatural base pair (UBP) is a designed subunit (or nucleobase) of DNA which is created in a laboratory and does not occur in nature. In 2012, a group of American scientists led by Floyd Romesberg, a chemical biologist at the Scripps Research Institute in San Diego, California, published that his team designed an unnatural base pair (UBP).[11] The two new artificial nucleotides or Unnatural Base Pair (UBP) were named d5SICS and dNaM. More technically, these artificial nucleotides bearing hydrophobic nucleobases, feature two fused aromatic rings that form a (d5SICS–dNaM) complex or base pair in DNA.[12][13] In 2014 the same team from the Scripps Research Institute reported that they synthesized a stretch of circular DNA known as a plasmid containing natural T-A and C-G base pairs along with the best-performing UBP Romesberg's laboratory had designed, and inserted it into cells of the common bacterium E. coli that successfully replicated the unnatural base pairs through multiple generations.[14] This is the first known example of a living organism passing along an expanded genetic code to subsequent generations.[12][15] This was in part achieved by the addition of a supportive algal gene that expresses a nucleotide triphosphate transporter which efficiently imports the triphosphates of both d5SICSTP and dNaMTP into E. coli bacteria.[12] Then, the natural bacterial replication pathways use them to accurately replicate the plasmid containing d5SICS–dNaM.
The successful incorporation of a third base pair is a significant breakthrough toward the goal of greatly expanding the number of amino acids which can be encoded by DNA, from the existing 20 amino acids to a theoretically possible 172, thereby expanding the potential for living organisms to produce novel proteins.[14] The artificial strings of DNA do not encode for anything yet, but scientists speculate they could be designed to manufacture new proteins which could have industrial or pharmaceutical uses.[16]
Length unit[edit]
Nucleotide (abbreviated "nt") is a common unit of length for single-stranded nucleic acids, similar to how base pair is a unit of length for double-stranded nucleic acids.
Nucleotide supplements[edit]
A study done by the Department of Sports Science at the University of Hull in Hull, UK has shown that nucleotides have significant impact on cortisol levels in saliva. Post exercise, the experimental nucleotide group had lower cortisol levels in their blood than the control or the placebo. Additionally, post supplement values of Immunoglobulin A were significantly higher than either the placebo or the control. The study concluded, "nucleotide supplementation blunts the response of the hormones associated with physiological stress."[17]
Another study conducted in 2013 looked at the impact nucleotide supplementation had on the immune system in athletes. In the study, all athletes were male and were highly skilled in taekwondo. Out of the twenty athletes tested, half received a placebo and half received 480 mg per day of nucleotide supplement. After thirty days, the study concluded that nucleotide supplementation may counteract the impairment of the body's immune function after heavy exercise.[18]
Abbreviation codes for degenerate bases[edit]
The IUPAC has designated the symbols for nucleotides.[19] Apart from the five (A, G, C, T/U) bases, often degenerate bases are used especially for designing PCR primers. These nucleotide codes are listed here. Some primer sequences may also include the character "I", which codes for the non-standard nucleotide inosine. Inosine occurs in tRNAs, and will pair with adenine, cytosine, or thymine. This character does not appear in the following table however, because it does not represent a degeneracy. While inosine can serve a similar function as the degeneracy "D", it is an actual nucleotide, rather than a representation of a mix of nucleotides that covers each possible pairing needed.
| Symbol[19] | Description | Bases represented | ||||
|---|---|---|---|---|---|---|
| A | adenine | A | 1 | |||
| C | cytosine | C | ||||
| G | guanine | G | ||||
| T | thymine | T | ||||
| U | uracil | U | ||||
| W | weak | A | T | 2 | ||
| S | strong | C | G | |||
| M | amino | A | C | |||
| K | keto | G | T | |||
| R | purine | A | G | |||
| Y | pyrimidine | C | T | |||
| B | not A (B comes after A) | C | G | T | 3 | |
| D | not C (D comes after C) | A | G | T | ||
| H | not G (H comes after G) | A | C | T | ||
| V | not T (V comes after T and U) | A | C | G | ||
| N | any base (not a gap) | A | C | G | T | 4 |




No comments:
Post a Comment